
Physiological determinants of urine and plasma myomiRNAs in recreational, middle-age athletes
Introduction
Micro RNAs (miRNAs), conventionally abbreviated as miRs or miRNAs, are short endogenous RNA molecules involved in a kaleidoscope of biological pathways (1). There is now consolidated evidence that miRNAs play a substantial role in many regulatory mechanisms, including cell development, differentiation, proliferation and apoptosis (2). A definite role of these short RNA molecules has hence been elucidated in relationship with several human pathologies, namely cancer and cardiovascular disease (2).
Growing interest is focusing on the potential diagnostic and prognostic applications of some highly muscle-specific miRNAs, typically called myomiRNAs (3), in the setting of skeletal muscle (4) and cardiac (5-7) injury. The former condition entails a variety of congenital disorders (e.g., muscular or myotonic dystrophies), along with acute muscle injuries due to direct muscular trauma, infections, toxic damage and exertional rhabdomyolysis. The potential clinical applications in the diagnostics of cardiomyocyte injuries principally include the diagnosis of myocardial ischemia and infarction, as well as the prediction or follow-up of cardiac hypertrophy and heart failure. The subset of muscle-specific miRNAs comprises a discrete number of molecules such as miRNA 1, miRNA 29-b, miRNA 133, miRNA 181-a, miRNA 206, miRNA 208 and miRNA 451. Among these, miRNA 133-a and miRNA 206 were found to be the most promising for understanding the biological response to physical activity and for the potential use for diagnosing muscle injury and in anti-doping testing (4,8).
Personalized or precision medicine is currently defined as an approach for disease treatment and prevention based on individual variability in genes, environment and lifestyle (9). The clinical use of whatever biomarkers should hence take into account the putative influence of many variables belonging to these domains, which most frequently include genetic factors, age, sex, ethnic origin, body composition, dietary habits, physical fitness and comorbidities (10). Therefore, the present study was aimed to identify the potential physiological determinants of miRNA 133-a and miRNA 206 in a population of recreational, middle-aged athletes.
Methods
Study population
The study group consisted of 28 healthy, middle-aged recreational athletes [11 females and 17 males; mean age: 46 years, age range 30–63 years; mean body mass index (BMI): 23.2 kg/m2, range: 18.5–27.9 kg/m2], belonging to a local team of amateur runners. Maximal oxygen consumption (VO2max) was measured by a running test on a treadmill through breath ergospirometric system (Quark B2, Cosmed, Rome, Italy). After appropriate familiarization, the athletes underwent a progressive incremental test, starting from habitual running pace and increasing the speed of 0.5 km/h every min until reaching volitional exhaustion.
All athletes were asked to stop training and to abstain from taking medications 48 hours before blood collection. The blood and urine for laboratory testing were collected early in the morning (i.e., between 8:30 and 9:30 AM) from subjects who had fasted for not less than 8 hours. Blood was drawn into evacuated blood collection tubes containing either K2 ethylenediaminetetraacetic acid (EDTA) or no additives (Terumo Europe N.V., Leuven, Belgium). A morning urine sample was also collected immediately after blood drawing. The blood samples were centrifuged at 1,300 ×g for 10 min at room temperature; serum and plasma EDTA were then separated and stored in aliquots at −70 °C until measurement. The urine samples were also centrifuged at 1,300 ×g for 10 min at room temperature and the supernatant was then divided in aliquots and stored at −70 °C until measurement.
Laboratory measurements
Total creatine kinase (CK) and cardiac troponin T (cTnT) were both measured in serum using a Roche Cobas 6,000 integrated analyzer (Roche Diagnostics GmbH, Mannheim, Germany). The concentration of cTnT was measured with a high-sensitivity (HS) immunoassay, displaying a limit of blank of 3.0 ng/L, a limit of detection of 5.0 ng/L and a 99th percentile upper reference limit (URL) of 14.0 ng/L (11). Total serum CK was also measured on Cobas 6,000, using the N-acetylcysteine-activated reference IFCC method.
The isolation of miRNA 133-a and miRNA 206 from plasma EDTA and urine samples was performed with the miRNeasy RNA isolation kit (Qiagen, Valencia, CA, USA) according to the manufacturer’s instructions. A standard volume of each plasma sample (200 μL) was supplemented with 5 mol of Caenorhabditis elegans miR-39 (Qiagen, Valencia, CA, USA) to normalize the results. miRNAs were reverse transcribed using the TaqMan MicroRNA Reverse Transcription Kit (Thermo Fisher Scientific, Australia) and the complementary DNA (cDNA) was used as template for the Real Time PCR. The reactions were performed on a 7,500 Real-Time PCR System (Applied Biosystems, CA, USA) using TaqMan MicroRNA assay (Applied Biosystems, Foster City, CA, USA). Briefly, qRT-PCR was carried out in a total volume of 20 μL containing 1.33 μL cDNA, 1× Universal PCR Master Mix and 1 μL gene-specific primers and probes. PCR parameters were as follows: 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s, and 60 °C for 1 min. The relative expression levels of miRNA 133-a and miRNA 206 were calculated by using the 2-∆Ct method (12).
Statistical analysis
Physiological and laboratory data were reported as median and interquartile range (IQR). The potential associations among the various parameters were evaluated using both univariate (Spearman’s correlation) and multivariate regression analyses. In multivariate regression analysis, the serum or urine concentration of myomiRNAs was entered as the dependent variable, whereas significant parameters in univariate analysis were entered as independent variables. Differences between groups were then tested with Mann Whitney U test. The statistical analysis was performed with Analyse-it (Analyse-it Software Ltd., Leeds, UK).
The study was part of an event called “Run For Science”, held in Verona (Italy) (the full information of the event can be retrieved from the Verona University Website, at: http://www.dsnm.univr.it/?ent=iniziativa&id=5382, Last accessed, 8 March 2017). The study was cleared by the local Institutional Review Board (Department of Neurological, Neuropsychological, Morphological and Movement Sciences, University of Verona), and was performed in accordance with the Declaration of Helsinki.
Results
The results of this study are shown in Tables 1 and 2. In univariate analysis, no single physiological or laboratory parameter was correlated with plasma miRNA 133-a concentration, whereas the urine concentration of miRNA 133-a was associated with both sex and BMI. Likewise, no single physiological or laboratory parameter was correlated with plasma miRNA 206 concentration, whereas the urine concentration of miRNA 206 was associated with both sex and serum cTnT. In multivariate logistic regression analysis, a modest association remained between urine miRNA 133-a and sex (beta coefficient, −9.9×10−6; P=0.046) but not with BMI (beta coefficient, −1.8×10−6; P=0.148). Similarly, urine miRNA 206 remained correlated with sex (beta coefficient, 2.4×10−5; P=0.037) but not with serum cTnT (beta coefficient, −1.4×10−6; P=0.510).
Table 1
| Variable | miRNA 133-a | |
|---|---|---|
| Plasma | Urine | |
| Age (years) | R= −0.23; P=0.230 | R= −0.14; P=0.483 |
| Sex | R= −0.03; P=0.873 | R= −0.59; P<0.001 |
| BMI (kg/m2) | R= −0.26; P=0.173 | R= −0.039; P=0.039 |
| VO2max (mL/min/kg) | R= 0.09; P=0.722 | R= 0.06; P=0.815 |
| Plasma miRNA 133-a | – | R= 0.05; P=0.784 |
| Urine miRNA 133-a | R= 0.05; P=0.784 | – |
| Plasma miRNA206 | R= 0.05; P=0.789 | R= 0.21; P=0.277 |
| Urine miRNA 206 | R= 0.12; P=0.545 | R= 0.05; P=0.784 |
| Serum CK (U/L) | R= 0.21; P=0.279 | R= −0.12; P=0.536 |
| Serum cTnT (ng/L) | R= −0.05; P=0.795 | R= −0.17; P=0.374 |
BMI, body mass index; VO2max, maximal oxygen consumption; CK, creatine kinase; cTnT, cardiac troponin T.
Table 2
| Variable | miRNA 206 | |
|---|---|---|
| Plasma | Urine | |
| Age (years) | R= −0.23; P=0.244 | R= −0.08; P=0.694 |
| Sex | R= −0.23; P=0.237 | R= 0.51; P=0.005 |
| BMI (kg/m2) | R= −0.15; P=0.446 | R= −0.24; P=0.217 |
| VO2max (mL/min/kg) | R= 0.02; P=0.940 | R= 0.11; P=0.673 |
| Plasma miRNA 133-a | R= 0.05; P=0.789 | R= 0.12; P=0.545 |
| Urine miRNA 133-a | R= 0.21; P=0.277 | – |
| Plasma miRNA206 | – | R= 0.30; P=0.124 |
| Urine miRNA 206 | R= 0.30; P=0.124 | R= 0.05; P=0.784 |
| Serum CK (U/L) | R= 0.09; P=0.656 | R= 0.16; P=0.408 |
| Serum cTnT (ng/L) | R= −0.28; P=0.148 | R= −038; P=0.046 |
BMI, body mass index; VO2max, maximal oxygen consumption; CK, creatine kinase; cTnT, cardiac troponin T.
The value distribution, differentiated between females and males of serum CK, serum cTnT, plasma and urine miRNA133-a and plasma and urine miRNA 206, is shown in Figure 1. In agreement with results of regression analyses, the urine values of miRNA133-a and miRNA 206 were found to be 3.0 and 1.9 folds higher in female than in male athletes. The Cohen’s d and the effect-size were 1.60 and 0.624 for miRNA 133-a, and 1.09 and 0.478 for miRNA 206, respectively. This implies that the percent overlap between the female and male populations of athletes was 27% for urine miRNA 133-a and 41% for urine miRNA 206, respectively. Notably, the Cohen’s d of the value distribution between female and male athletes was 0.035 for plasma miRNA 133-a and 0.356 for plasma miRNA 206, so highlighting a very large percent overlap of values between sexes (i.e., ~97% for miRNA 133-a and ~76% for miRNA 206, respectively).
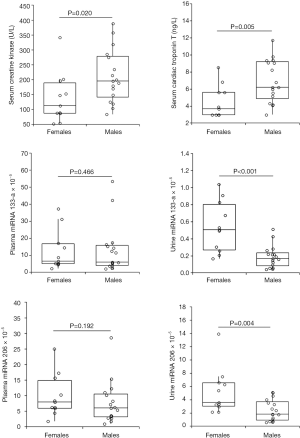
A significant difference was also observed for CK and cTnT, with the serum values of both biomarkers being nearly 40% higher in male than in female athletes. In no case did the concentration of serum cTnT exceeded the diagnostic cut-off of cardiac injury (i.e., the 99th percentile URL; 14.0 ng/L). The Cohen's d and the effect-size were 0.87 and 0.399 for serum CK, and 1.22 and 0.521 for serum cTnT, respectively. This implies that the percent overlap between the female and male populations of athletes was 48% for CK and 38% for cTnT, respectively.
Discussion
Several lines of evidence now attest that miRNAs are active players in health and disease (13-15). Taken together, our results suggest that the baseline plasma and urine concentration of the two main myomiRNA measured in this study is virtually independent from age, BMI, physical fitness, (i.e., VO2max), as well as from skeletal (i.e., serum CK) and cardiac (i.e., serum cTnT) muscle health. Although the plasma values of both miRNA 133-a and miRNA 206 were also found to be independent from sex, the urine concentration of these two myomiRNA was considerably higher in female than in male athletes. This is a rather unexpected finding, since the serum values of the two more conventional biomarkers of skeletal muscle (i.e., CK) and myocardiocyte (i.e., cTnT) injury were instead increased in male compared to female athletes (Figure 1).
The evidence that the biological pathways underlying the metabolism of myomiRNA and muscle injury biomarkers in resting subjects is quite divergent, and is also totally independent from the overall BMI, paves the way to some important reflections. First, the lack of correlation between plasma and urine concentration of both miRNA 133-a and 206 implies that the measurement of these myomiRNAs in different body fluids is not interchangeable. No previous studies have assessed miRNA 133-a or miRNA 206 in urine samples of middle aged healthy subjects to the best of our knowledge, so this finding is highly innovative and underscores that increased renal clearance, and/or reduced reabsorption, may play a significant role in myomiRNAs metabolism. Then, the strong and independent association that we have observed between sex and urine concentration of both miRNA 133-a and miRNA 206 also implies that the urine reference ranges of these biomarkers should be partitioned for the sex. This is especially true for miRNA133-a, since not only was its urine concentration was 3-fold higher in women than in men, but also the percent overlap between the values of male and female athletes was extremely modest (i.e., 27%). Despite the overlap between values in men and women was less accentuated for miRNA 206 (i.e., 41%), the 1.9-fold increase of urine value of this myomiRNA suggests that its reference range should also be partitioned for the sex. Another substantial aspect highlighted by our data is that case-control studies involving the measurement of myomiRNAs in urine should be planned to exactly match the different study populations for the sex. This is especially important if one considers that miRNAs have a good stability in urine and under many storage conditions, so supporting their increasing use in urine as diagnostic biomarkers for several pathologies (16).
Regardless of these general considerations, it remains unclear why the urine concentration of myomiRNAs should be higher in women than in men, especially if one considers that their plasma concentration is virtually identical between sexes (Figure 1). The potential influence of renal function (i.e., glomerular filtration rate) on the clearance of plasma miRNA 133a has been earlier shown (17), but this is not helpful to explain the different urine concentration between women and men observed in our investigation. This aspect should probably be targeted by future studies focusing on clearance of myomiRNAs in humans.
Conclusions
In conclusion the results of this study demonstrate that the plasma concentration of miRNA 133-a and 206 is virtually independent from the most common physiological variables, whereas the urine reference values of these two myomiRNAs should be partitioned for the sex. As regards the future perspectives, the results of our study lead the way to an exciting scenario, where myomiRNA measurement may be used for creating individual training plans for athletes (i.e., personalized training).
Acknowledgments
Fabian Sanchis-Gomar is supported by a post-doctoral contract (APOSTD/2016/140) granted by the Regional Ministry of Education, Research, Culture and Sport (Valencia).
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/jlpm.2017.04.04). Fabian Sanchis-Gomar serves as an unpaid editorial board member of Journal of Laboratory and Precision Medicine from December 2016 to November 2018. Martina Montagnana serves as the unpaid Associate Editor-in-Chief of Journal of Laboratory and Precision Medicine from November 2016 to October 2021. Giuseppe Lippi serves as the unpaid Editor-in-Chief of Journal of Laboratory and Precision Medicine from November 2016 to October 2021. The authors have no other conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was cleared by the local Institutional Review Board (Department of Neurological, Neuropsychological, Morphological and Movement Sciences, University of Verona, Verona, Italy). Individual informed consent was waived.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004;116:281-97. [Crossref] [PubMed]
- Cai Y, Yu X, Hu S, et al. A brief review on the mechanisms of miRNA regulation. Genomics Proteomics Bioinformatics 2009;7:147-54. [Crossref] [PubMed]
- Malizia AP, Wang DZ. MicroRNAs in cardiomyocyte development. Wiley Interdiscip Rev Syst Biol Med 2011;3:183-90. [Crossref] [PubMed]
- Zacharewicz E, Lamon S, Russell AP. MicroRNAs in skeletal muscle and their regulation with exercise, ageing, and disease. Front Physiol 2013;4:266. [Crossref] [PubMed]
- Lippi G, Mattiuzzi C, Cervellin G. Circulating microRNAs (miRs) for diagnosing acute myocardial infarction: meta-analysis of available studies. Int J Cardiol 2013;167:277-8. [Crossref] [PubMed]
- Klug G, Metzler B. The role of circulating microRNAs in acute coronary syndromes: ready for prime time? Ann Transl Med 2016;4:537. [Crossref] [PubMed]
- Adam R, Kelly D. Is there a role for microRNAs as novel predictors of prognosis in myocardial infarction? Ann Transl Med 2016;4:473. [Crossref] [PubMed]
- Lombardi G, Perego S, Sansoni V, et al. Circulating miRNA as fine regulators of the physiological responses to physical activity: Pre-analytical warnings for a novel class of biomarkers. Clin Biochem 2016;49:1331-9. [Crossref] [PubMed]
- Lippi G, Montagnana M. Laboratory and precision medicine: the reasons for the birth of a new journal. J Lab Precis Med 2016;1:3. [Crossref]
- Lippi G, Bassi A, Bovo C. The future of laboratory medicine in the era of precision medicine. J Lab Precis Med 2016;1:7. [Crossref]
- Giannitsis E, Kurz K, Hallermayer K, et al. Analytical validation of a high-sensitivity cardiac troponin T assay. Clin Chem 2010;56:254-61. [Crossref] [PubMed]
- Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta C(T)) method. Methods 2001;25:402-8. [Crossref] [PubMed]
- Yu H, Lu Y, Li Z, et al. microRNA-133: expression, function and therapeutic potential in muscle diseases and cancer. Curr Drug Targets 2014;15:817-28. [Crossref] [PubMed]
- McCarthy JJ. MicroRNA-206: the skeletal muscle-specific myomiR. Biochim Biophys Acta 2008;1779:682-91. [Crossref] [PubMed]
- Lippi G, Mattiuzzi C. The biomarker paradigm: between diagnostic efficiency and clinical efficacy. Pol Arch Med Wewn 2015;125:282-8. [PubMed]
- Mall C, Rocke DM, Durbin-Johnson B, et al. Stability of miRNA in human urine supports its biomarker potential. Biomark Med 2013;7:623-31. [Crossref] [PubMed]
- Gidlöf O, Andersson P, van der Pals J, et al. Cardiospecific microRNA plasma levels correlate with troponin and cardiac function in patients with ST elevation myocardial infarction, are selectively dependent on renal elimination, and can be detected in urine samples. Cardiology 2011;118:217-26. [Crossref] [PubMed]
Cite this article as: Danese E, Benati M, Sanchis-Gomar F, Tarperi C, Salvagno GL, Paviati E, Montagnana M, Schena F, Lippi G. Physiological determinants of urine and plasma myomiRNAs in recreational, middle-age athletes. J Lab Precis Med 2017;1:13.


