
Methylation panel as a diagnostic biomarker in Barrett’s oesophagus: a comprehensive biomarker panel in a population-based screening programme?
We congratulate the authors for their excellent paper and contribution to the body of research within diagnostic biomarkers for Barrett’s oesophagus (BO). This work is a promising step towards the development of improved methods of screening for BO and the earlier diagnosis of oesophageal adenocarcinoma (OAC).
In the UK and USA screening for BO is reserved for those patients with symptoms of chronic gastroesophageal reflux disease (GORD) and with specific risk factors. Currently, diagnosis of BO is reliant upon endoscopy; an expensive and invasive procedure which requires technical skill, making its adoption into a population screening programme difficult. This presents a dilemma; the majority of cases of BO are undiagnosed, and despite a relatively low rate conversion rate to OAC (probably around 0.3% per year), the incidence of both BO and OAC is rising (1,2). The cause of this rise is not due to limitations in curative endoscopic therapy. On the contrary, developments in recent years have delivered significant advances in endoscopic treatment. Techniques such as radiofrequency ablation, endoscopic mucosal resection and endoscopic submucosal resection have all provided more opportunities for curative therapy with lower complication rates than surgical intervention (3).
In cancer timely diagnosis improves outcome and this is particularly true of OAC. Mortality rates from this cancer remain over 80% at 5 years unless detected early (4), highlighting the need to develop more feasible ways to screen larger populations and ultimately, make diagnoses earlier. In the search for more suitable screening methods, alternatives to traditional endoscopy have been sought with varying success. Nasal endoscopy and string capsule video endoscopy have been trialled but are expensive and require significant expertise. Non-endoscopic screening tests such as the “Cytosponge” have been developed and it is this method, which involves a patient swallowing a sponge within a soluble capsule attached to a thin string, that is utilised in Chettouh et al.’s 2017 study (Figure 1) (5). This technique, which is inexpensive, minimally invasive and acceptable to patients, seems to provide a viable alternative to traditional endoscopic methods (6). Furthermore, it seems suitable for the primary care setting and may bridge the gap from endoscopy suite to GP consultation room. This approach is able to reach a much wider group of patients with gastro oesophageal reflux disease at risk of developing BO and cancer.
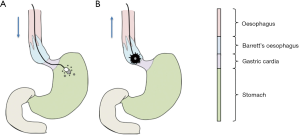
The principle of the Cytosponge is cell retrieval from the lining of the oesophagus. Besides cell retrieval, success of the system requires that immunohistochemical techniques to differentiate BO and OAC from normal squamous epithelium and gastric cardia. Biomarkers including TFF3 are one such defining feature and have a sensitivity and specificity of 79.9% and 92.4% respectively for BO (7). Although a promising start, difficult specimen processing requirements and a relatively low sensitivity suggest that other biomarkers should be sought. One such biomarker is DNA methylation. The methylation of DNA is an epigenetic change that is found in a number of different cancers. In particular, it seems that the change is an important step in the development and progression of BO as evidenced by studies which demonstrate this precise mutation in genes found in BO.
In Chettouh et al.’s study, methylated genes were identified from samples gathered in the BEST2 trial initially designed to assess the viability of the TFF3 biomarker in BO using the Cytosponge. Using these tissues, pilot and validation studies demonstrated a total of 10 genes which were hyper-methylated in Barrett’s cells. Four of these (TFP12, TWIST1, ZNF345 and ZNF569) were highly differentially methylated and two were novel (ZNF345 and ZNF569), having never previously been shown to be hyper-methylated in cancer. Encouragingly, sensitivity and specificities were comparable to those seen in BEST2 and the tissues in which genes were not hyper-methylated tended to have smaller circumferential (C) and maximal (m) segments, in keeping with studies that have shown that longer segments are more likely to progress to OAC (8). We have provided Table 1 as a summary of the relative strengths and weaknesses of this approach.
Table 1
| Chettouh et al.’s study | Requires prospective studies |
|---|---|
| Strengths | |
| ❖ Large validation study | Yes |
| ❖ Differentiation between BO and gastric cardia | Yes |
| ❖ Sensitivity and specific of four most differentially methylated genes is comparable to BEST2 | – |
| Weaknesses | |
| ❖ Retrospective analysis | Yes |
| ❖ Inability to differentiate between NDBO and DBO | Yes |
| ❖ Likely to be used within biomarker panel (rather than in isolation) for a population-based screening programme | – |
NDBO, non-dysplastic Barrett’s oesophagus; DBO, dysplastic Barrett’s oesophagus.
This research is a retrospective analysis utilising samples from the BEST2 trial and it is clear that prospective validation is required to develop the progress made. Before the development of a population-based screening programme, a number of important questions need to be answered. In particular, how will the technology be integrated into a screening program and how comprehensive a biomarker panel is required to reach a sufficient level of sensitivity and specificity? Furthermore, will a screening method be required to identify dysplastic lesions as well as non-dysplastic BO (NDBO)? Within a screening program that is unable to detect dysplasia it is likely that patient risk stratification would need to carefully considered to ensure malignancy is not missed in those patients at highest risk of developing OAC.
Although this study highlights four particularly hyper-methylated genes, a number of others were identified and there is clearly a diverse profile of genetic changes which may lead to the development of BO. As identified in this study, DNA methylation of specific genes is likely to contribute significantly to the development of BO. However, for a population based screening programme to be successful, with whatever the cell retrieval process, it is becoming increasingly clear that identification of this pre-malignant condition is likely to include a number of genes across a comprehensive biomarker panel.
Acknowledgments
Funding: None.
Footnote
Provenance and Peer Review: This article was commissioned and reviewed by the Section Editor Dr. Dao-Jun Hu (Department of Clinical Laboratory, Xin Hua Hospital Affiliated to Shanghai Jiao Tong University School of Medicine, Chongming Branch, Shanghai, China).
Conflicts of Interest: Both authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/jlpm.2018.04.01). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Vaughan TL, Fitzgerald RC. Precision prevention of oesophageal adenocarcinoma. Nat Rev Gastroenterol Hepatol 2015;12:243-8. [Crossref] [PubMed]
- Lochhead P, Chan AT. Screening and surveillance for Barrett esophagus. JAMA Intern Med 2015;175:159-60. [Crossref] [PubMed]
- Pimentel-Nunes P, Dinis-Ribeiro M, Ponchon T, et al. Endoscopic submucosal dissection: European society of gastrointestinal endoscopy (ESGE) guideline. Endoscopy 2015;47:829-54. [Crossref] [PubMed]
- Medical Research Council Oesophageal Cancer Working Group. Surgical resection with or without preoperative chemotherapy in oesophageal cancer: a randomised controlled trial. Lancet 2002;359:1727-33. [Crossref] [PubMed]
- Chettouh H, Mowforth O, Galeano-Dalmau N, et al. Methylation panel is a diagnostic biomarker for Barrett’s oesophagus in endoscopic biopsies and non-endoscopic cytology specimens. Gut 2017; [Epub ahead of print]. [Crossref] [PubMed]
- Kadri SR, Lao-Sirieix P, O’Donovan M, et al. Acceptability and accuracy of a non-endoscopic screening test for Barrett’s oesophagus in primary care: cohort study. BMJ 2010;341:c4372. [Crossref] [PubMed]
- Ross-Innes CS, Debiram-Beecham I, O’Donovan M, et al. Evaluation of a minimally invasive cell sampling device coupled with assessment of trefoil factor 3 expression for diagnosing Barrett’s esophagus: a multi-center case-control study. PLoS Med 2015;12:e1001780 [Crossref] [PubMed]
- Wani S, Falk GW, Post J, et al. Risk factors for progression of low-grade dysplasia in patients with Barrett’s esophagus. Gastroenterology 2011;141:1179-86. [Crossref] [PubMed]
Cite this article as: Riley T, Ang Y. Methylation panel as a diagnostic biomarker in Barrett’s oesophagus: a comprehensive biomarker panel in a population-based screening programme? J Lab Precis Med 2018;3:37.


