
Detection of 8-OHdG as a diagnostic biomarker
It is widely documented that the accumulation of reactive oxygen species (ROS) can result in structural cellular and/or genetic changes modulating essential triggers of tumor formation, especially at the initial steps of carcinogenesis. In nuclear and mitochondrial DNA, 8-hydroxy-2’-deoxyguanosine (8-OHdG) or 8-oxo-7,8-dihydro-2’-deoxyguanosine (8-oxodG) are the most commonly observed single nucleotide-base lesions that might induce mutations in replicating DNA. Also, it is well accepted that these free radical-induced oxidative lesions are potential biomarkers of oxidative DNA damage (1,2), though continuously repaired by base excision repair (BER) in cells.
In mammalian cells, 8-OHdG is repaired by a dedicated enzyme, 8-oxoguanine DNA glycosylase-1 (OGG1), which is one of the major players of BER. When the oxidized base 8-OHdG is not removed by OGG1 via glycosylation from the DNA backbone, the accumulation of damaged bases may lead to transversion mutations of G to T resulting in alterations of the genomic context. Eventually, this may trigger or exacerbate diverse pathological conditions (3,4).
Evaluation of 8-OHdG damage as a diagnostic biomarker
The generation and the accumulation of ROS might lead to oxidative damage in the DNA which in turn could bring about DNA/protein-related pathologies. Thus, the biological significance of alterations in 8-OHdG levels has been studied in a wide range of diseases. A stimulatory role of oxidative damage in inflammation has been shown for inflammatory bowel disease (5) and type 2 diabetes (6). Also, elevated levels of 8-OHdG have been reported to be correlated with clinical outcomes of stroke and have a close relationship with atherosclerotic plaque types and vascular recurrence in non-cardio embolic stroke patients (7-9).
In addition, when serum 8-OHdG values in healthy controls and patients suffering from nonalcoholic fatty liver (NAFL) and nonalcoholic steatohepatitis (NASH) were compared, the serum 8-OHdG was identified as a diagnostic marker for NASH (10). An increased serum level of 8-oxoG with a concomitantly decreased OGG1 protein expression was shown in Alzheimer’s disease patients (11). Thus, 8-oxoG was suggested as a blood biomarker for Alzheimer’s disease that can assist in the stratification of these patients regarding treatment options (11).
Furthermore, oxidative damage seems to play an active role in aging and the use of antioxidants might be a straightforward anti-aging strategy option (12-14). Thus, not surprisingly, 8-OHdG has been investigated as a potential marker of age-related damage accumulation in several studies (12,13,15). Moreover, increased 8-oxoG levels have been reported in animal studies using tissue and urine samples (12,13). This adds further evidence to the assumption that oxidative base damage could be a characteristic feature of aging. In order to elucidate the age-dependent alterations in humans, 8-OHdG damage levels have been investigated in different age groups and an age-dependent increase in oxidized guanine in both DNA and RNA by comparing 21- to 30-year-old individuals to 81- to 90-year-old ones regarding both genders have been shown (15). These data suggested 8-oxo-7,8-dihydro-2’-deoxyguanosine (8-oxodGsn) and 8-oxo-7,8-dihydroguanosine (8-oxoGsn) as promising aging biomarkers that could be utilized for determining age-related pathologies (15).
Of note, increased 8-OHdG levels are closely related to the accumulation of mutations triggering carcinogenesis. In a meta-analysis, an enzyme-linked immunosorbent assay (ELISA)-based method was used to measure the 8-OHdG concentration in leukocyte DNA of 314 patients with serous ovarian cancer (SOC) and 774 normal controls (16). Interestingly, a high 8-OHdG concentration in leukocyte DNA was associated with advanced age and poor prognosis of SOC patients in comparison to controls (16). In addition, increased 8-OHdG levels were found as a triggering factor for inflammation-related carcinogenesis (17).
In another report, 8-oxodG was studied as an indicator of oxidative DNA damage in patients with coronary artery disease (CAD) before and after coronary artery bypass grafting (CABG). Urinary 8-oxodG levels were detected by a liquid chromatography-coupled tandem mass spectrometer (LC-MS/MS) method and results indicated higher 8-oxodG levels in CAD patients than in controls. The ascertained short-term oxidative DNA damage was induced by the CABG procedure in the CAD patients whereas complete recovery occurred six months later after the surgical operation (18). Furthermore, to investigate the association between 8-OHdG levels and heart failure, a comprehensive meta-analysis elicited that 8-OHdG levels are higher in patients with heart failure than in controls (19).
The use of 8-OHdG has also been found beneficial for the assessment of exercise-induced oxidative damage in several studies (20-23). Although most of the studies have not concluded a solid link between exercise and oxidative damage, there is a tendency of increased 8-OHdG levels during extensive exercise. Consistently, the level of urinary 8-OHdG was higher in physically active subjects undergoing extensive exercise, when oxidative response markers were compared in active middle-aged subjects with those in sedentary individuals (24). However, regular exercise has even health benefits partly because it reportedly lowers the levels of oxidation products of proteins and DNA at rest and also most likely due to the increased stress tolerance via activated DNA damage mechanisms (25). To gain a detailed insight into the physiological mechanisms involved under extreme conditions, a group of experienced ultra-marathon runners was monitored (26). According to this study, exhaustive and prolonged exercise not only promotes the generation of ROS but also induces oxidative stress, temporary renal weakening as well as inflammation (26). Vigorous exercise amounting to approximately 10 h a day for 30 days increased the rate of oxidative DNA modifications by 33% in 20 men owing to the urinary excretion of 8-OHdG (27).
In contrast, there are several studies based on the analysis of urinary excretion of 8-OHdG claiming that no significant accumulation of oxidative DNA damage is caused by various exercise types such as repeated bouts of cycling (20), long-distance running (28) and swimming (29). In another comprehensive study correlating gender and lifestyle factors such as diet and exercise, the authors observed no significant differences between groups of trained and untrained men and women when protein/lipid/DNA oxidative stress markers including 8-OHdG were analyzed (30).
Furthermore, in a third group of studies with repeated exercise in long distance runners, oxidative DNA damage was found intensely increased when urine samples were evaluated for 8-OHdG levels (31-33). The data revealed that oxidative damage to DNA is not accumulated in the long term possibly because of an adaptation in damage repair capacity, leading to normalization of DNA damage. A similar conclusion was made by another study investigating urinary 8-OHdG levels of eight professional cyclists after long-term exercises. The results demonstrated that cycling induced oxidative DNA damage, which was sustained as long as the exercise was repeated with subsequent adaptation of antioxidant defense, leading the normalization in 8-OHdG excretion (34).
Methods of 8-OHdG detection
Quantitation of 8-OHdG can be carried out with a wide range of methodologies currently. Several of them use different antibody labeling techniques to allow the detection of 8-OHdG such as ELISA, immunohistochemistry (IHC) technique or immunofluorescence (IF) imaging. However, some of these methods require DNA isolation for 8-OHdG analysis at first. The commonly used LC-MS/MS is one of them and exhibits significant advantages by producing highly precise and sensitive data at femtomolar levels of genomic DNA from various sample materials including tissue, plasma and urine (12,13,35,36). Altogether, a pool of extracted DNA from various sources can be used for the absolute quantitation of 8-OHdG by several techniques. For comparative analysis, the ratio of 8-OHdG/dG needs to be determined. Last but not least, this requires a standard curve covering the measurement ranges of both modified and dG bases which represents a major disadvantage of this approach. Thus, respective ratios of 1/105 were reported in control genomic DNA. Therefore, the analysis of the 8-OHdG/dG ratio and their absolute quantities by LC-MS/MS necessitates a dynamic range of linearity of the corresponding standard curve of more than 6 logs. Another common disadvantage of the LC-MS/MS analysis relates to the quantitative determination of total cellular oxidative damage due to the use of pooled DNA. This is a technical limitation and derives from the method itself.
Of note, high-performance liquid chromatography (HPLC) is another commonly preferred alternative method for the precise quantitation of 8-OHdG in various sample types (20,37) with similar advantages and disadvantages like LC-MS/MS.
A readily available assay technique used in many studies for the quantitation of 8-OHdG damage is the ELISA (11,36,38). Likewise in MS/MS analysis, the method requires DNA isolation at the beginning, digestion thereof and hybridization subsequently. Although, there are several commercial kits available, the major limitation of this method lies in its accuracy and precision; a specific antibody is also required to perform the assay appropriately.
Other assay techniques such as IHC (5,17), immunocytochemistry (ICC) (39) and IF (40-43) have been used to develop cell/tissue-specific assays too. However, these assays are commonly characterized by imprecise and semi-quantitative data acquisition demanding complex signal density analysis with extensive operator time. Thus, they have been employed mostly in conceptual studies. The read-out is usually given as an average signal intensity. Furthermore, these methods have to contend with background noise, since cytoplasmic RNA staining interferes with the DNA-specific quantitation of 8-OHdG.
Automated IF-image analysis employing sophisticated pattern recognition may be a remedy for these challenges. Recently, the IF automated interpretation system Aklides system was used for quantitation of 8-OHdG by analyzing fluorescent 8-OHdG foci within the nuclei of cells with oxidative damage (44). This interpretation system enables the pattern recognition of fluorescent foci and, thus, permits the quantitation of their fluorescent signals with regard to their shape and size. Of note, the Aklides system was optimized for quantitative analysis of DNA double strand break γ-H2AX(S139) foci analysis originally (45,46). Recent studies have demonstrated the efficiency of this automated system for DNA damage providing the opportunity to run studies with large samples numbers (47-50). It can be utilized for clinical purposes to ascertain drug resistance and radiotherapy sensitivity in patients (51-53). The method allows to measure DNA damage levels at a single focus level.
Our group could demonstrate consistently that oxidative damage can be ascertained by foci formation in nuclei of damaged cells (44). This has been rendered possible by a modified IF-labeling procedure allowing to analyze foci occurrence in single nuclei as well as the size of these foci. Since the γ-H2AX(S139) foci formations appears to be similar to the one of 8-OHdG foci, automated IF interpretation could be used readily for the comprehensive analysis of 8-OHdG foci (Figure 1). Thus, 8-OHdG occurrence with subsequent damage quantitation has been performed to compare results obtained by the Aklides interpretation system with LC-MS/MS data (44).
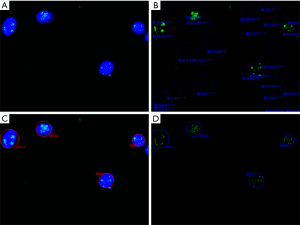
Altogether, in order to choose the most appropriate method for 8-OHdG quantitation, the advantages and disadvantages of each method must be comprehensively evaluated. Although the total amount of 8-OHdG in various samples such as urine, plasma, and tissue can be measured accurately, two significant limitations should be considered. Firstly, the basal amount of the damaged 8-OHdG is usually quite small to be assessed accurately with analytical methods, as the values of the damaged bases can be very close to the detection limits of the methods used. Secondly, damage measured across the entire genomic DNA can lead to misleading results, as most of the damage detected comes from non-coding regions of the genome. As a fact, most DNA is not encoding and the induction and rates of DNA repair in coding and non-coding regions could be very different (4). As an alternative method for measuring the total damage, ELISA has the disadvantage that the detection limit cannot reach the individual value of LC-MS/MS. Since the DNA isolation step carried out in analytical methods such as ELISA can misinterpret the total damage level due to the induction of additional DNA oxidation during purification, IF-based methods without DNA isolation should be considered. Another advantage is the shorter sample preparation time. A study comparing urine levels of 8-OHdG in healthy individuals with LC-MS/MS, HPLC-EC and ELISA found a significant discrepancy between chromatography and immunoassay approaches, demonstrating the importance of proper method selection with appropriate detection limits (54).
In addition, the most favorable body sample should be selected for biologically relevant results, as blood levels may not directly represent the level of damage in the cells or tissues exposed to DNA-damaging contaminants (4). A recent study comparing plasma and urine as sample materials demonstrated that plasma measurements were more sensitive to training-induced 8-OHdG changes than urine measurements regarding the reliable measurement of 8-OHdG damage (38).
In contrast, IF methods have certain benefits such as being performed with low amounts/numbers of samples/cells in comparison to the analytical methods, in which more cells or material is required to isolate relatively higher amounts of DNA for reliable quantitation of the damaged base. Since analysis of DNA damage in a single cell is necessary to investigate genomic heterogeneity and related specific damage formation and corresponding repair responses in carcinogenesis studies, IF-based techniques present advantages in quantitation of potentially heterogeneous damage levels among cells from both blood and tissue. The automated interpretation by such automated IF systems like the Aklides can provide the cell-based quantitative analysis of 8-OHdG in any cell type after cell-specific optimization of computational parameters for pattern recognition.
Conclusions
It is generally accepted that 8-OHdG or 8-oxodG are potential biomarkers for measuring the effect of endogenous oxidative damage to DNA as an important factor of initiation and promotion of carcinogenesis. These biomarkers have been used to assess DNA damage in humans after exposure to various causes of cancer such as oxidant chemicals, heavy metals, smoke and polycyclic aromatic hydrocarbons. In addition, 8-OHdG has been used in many studies in recent years not only as a biomarker for the measurement of endogenous oxidative DNA damage but also to ascertain the risk for diseases development and intensified aging. Overall, the studies showed that urinary 8-OHdG is a good biomarker for risk assessment of various cancers and degenerative diseases. Although some of the most widely used methods enable quantitative analysis such as HPLC with electrochemical detection and/or tandem mass spectrometry, these techniques are characterized by various advantages and disadvantages (Table 1). In order to overcome the methodological problems of 8-OHdG analysis, the European Standards Committee (ESC) for Oxidative DNA Damage summarized the artefactual oxidation problems faced during isolation and purification of oxidative DNA products in 1997. Automated IF-interpretation systems provide fast and reliable 8-OHdG analysis which may complement the ESC standards for comprehensive 8-OHdG foci testing. They offer a cell-based quantitation of individual oxidative damage that can be applied to several sample materials and thus might offer new opportunities in precision medicine.
Table 1
| Method for 8-OHdG detection | Advantages | Disadvantages | LOD/LOQ | Matrix | References |
|---|---|---|---|---|---|
| LC-MS/MS | Highly precise and sensitive assessment at femtomolar levels of genomic DNA | Requirement of DNA isolation leading to additional DNA oxidation during purification. Quantification of 8-OHdG/dG levels needs a standard curve covering both measurement ranges with a dynamic range of linearity of more than 6 logs. Quantitation of total cellular oxidative damage (no single cell-based data) | Despite non-linearity at low concentrations, a LOD of 0.3 nM and a LOQ of 1.0 nM are reported | Tissue, plasma and urine | (12,13,35,36) |
| UHPLC | Quantitative data acquisition | Similar to LC-MS/MS | Despite non-linearity at low concentrations, a LOD of 0.3 nM and a LOQ of 1.0 nM are reported | Tissue, plasma and urine | (20,37) |
| IHC/ICC/IF | Shorter sample preparation time Low sample amount for detection in comparison to analytical methods | Semi-quantitative data acquisition. Interference of the DNA-specific quantitation of 8-OHdG by staining of cytoplasmic RNA | N/A* | Cell culture, tissue | IHC (5,17); ICC (39); IF (40-43) |
| ELISA | Availability of commercial kits | Requirement of DNA isolation leading to additional DNA oxidation during purification. Limited precision due to antibody dependency | 2 pg of 8-OHdG base of isolated DNA | Cell culture, tissue, plasma and urine | (11,36,38) |
| AKLIDES | Cell-based data to study genomic heterogeneity. Quantitation of DNA damage without interference of damaged RNA. Shorter sample preparation time in comparison to analytical methods. Low sample amount for detection in comparison to analytical methods | Protocol optimization and adjustment of system parameters is required for different cell types | An 8-OHdG focus represents a single damaged base in 5 pg of genomic DNA | Cell culture, PBMC/plasma | 44 |
*, not applicable to these assays. ELISA, enzyme-linked immunosorbent assay; ICC, immunocytochemistry; IF, indirect immunofluorescence; IHC, immunohistochemistry; LC-MS/MS, liquid chromatography-mass spectrometry/mass spectrometry; LOD, limit of detection; LOQ, limit of quantification; PBMC, peripheral blood mononuclear cell; UHPLC, ultra-high performance liquid chromatography.
Acknowledgments
Funding: This research was supported by a grant (113S044) from Turkish Scientific and Technology Research Council (TUBITAK) to KS Korkmaz.
Footnote
Provenance and Peer Review: This article was commissioned by the editorial office, Journal of Laboratory and Precision Medicine for the series “DNA Damage Assessment for Precision Medicine”. The article has undergone external peer review.
Conflicts of Interest: The series “DNA Damage Assessment for Precision Medicine” was commissioned by the editorial office without any funding or sponsorship. Dirk Roggenbuck served as an unpaid Guest Editor of the series and serves as an unpaid editorial board member of Journal of Laboratory and Precision Medicine from August 2017 to July 2019. D Roggenbuck is one of the shareholders of Medipan GmbH; GA Generic Assays GmbH being diagnostic manufacturers. The authors have no other conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Kryston TB, Georgiev AB, Pissis P, et al. Role of oxidative stress and DNA damage in human carcinogenesis. Mutat Res 2011;711:193-201. [Crossref] [PubMed]
- Cadet J, Loft S, Olinski R, et al. Biologically relevant oxidants and terminology, classification and nomenclature of oxidatively generated damage to nucleobases and 2-deoxyribose in nucleic acids. Free Radic Res 2012;46:367-81. [Crossref] [PubMed]
- Ba X, Aguilera-Aguirre L, Rashid QT, et al. The role of 8-oxoguanine DNA glycosylase-1 in inflammation. Int J Mol Sci 2014;15:16975-97. [Crossref] [PubMed]
- Paz-Elizur T, Sevilya Z, Leitner-Dagan Y, et al. DNA repair of oxidative DNA damage in human carcinogenesis: potential application for cancer risk assessment and prevention. Cancer Lett 2008;266:60-72. [Crossref] [PubMed]
- Ding X, Hiraku Y, Ma N, et al. Inducible nitric oxide synthase-dependent DNA damage in mouse model of inflammatory bowel disease. Cancer Sci 2005;96:157-63. [Crossref] [PubMed]
- Nakamura A, Osonoi T, Terauchi Y. Relationship between urinary sodium excretion and pioglitazone-induced edema. J Diabetes Investig 2010;1:208-11. [Crossref] [PubMed]
- Hsieh YW, Lin KC, Korivi M, et al. The reliability and predictive ability of a biomarker of oxidative DNA damage on functional outcomes after stroke rehabilitation. Int J Mol Sci 2014;15:6504-16. [Crossref] [PubMed]
- Brea D, Roquer J, Serena J, et al. Oxidative stress markers are associated to vascular recurrence in non-cardioembolic stroke patients non-treated with statins. BMC Neurol 2012;12:65. [Crossref] [PubMed]
- Chen YC, Chen CM, Liu JL, et al. Oxidative markers in spontaneous intracerebral hemorrhage: leukocyte 8-hydroxy-2'-deoxyguanosine as an independent predictor of the 30-day outcome. J Neurosurg 2011;115:1184-90. [Crossref] [PubMed]
- Jiang Y, Han T, Zhang ZG, et al. Zhonghua Nei Ke Za Zhi 2017;56:34-8. [The significance of 8-hydroxy-deoxyguanosine acid in the diagnosis of nonalcoholic steatohepatitis]. [PubMed]
- Sliwinska A, Kwiatkowski D, Czarny P, et al. The levels of 7,8-dihydrodeoxyguanosine (8-oxoG) and 8-oxoguanine DNA glycosylase 1 (OGG1) - A potential diagnostic biomarkers of Alzheimer's disease. J Neurol Sci 2016;368:155-9. [Crossref] [PubMed]
- Gan W, Nie B, Shi F, et al. Age-dependent increases in the oxidative damage of DNA, RNA, and their metabolites in normal and senescence-accelerated mice analyzed by LC-MS/MS: urinary 8-oxoguanosine as a novel biomarker of aging. Free Radic Biol Med 2012;52:1700-7. [Crossref] [PubMed]
- Nie B, Gan W, Shi F, et al. Age-dependent accumulation of 8-oxoguanine in the DNA and RNA in various rat tissues. Oxid Med Cell Longev 2013;2013:303181 [Crossref] [PubMed]
- Simioni C, Zauli G, Martelli AM, et al. Oxidative stress: role of physical exercise and antioxidant nutraceuticals in adulthood and aging. Oncotarget 2018;9:17181-98. [Crossref] [PubMed]
- Gan W, Liu XL, Yu T, et al. Urinary 8-oxo-7,8-dihydroguanosine as a Potential Biomarker of Aging. Front Aging Neurosci 2018;10:34. [Crossref] [PubMed]
- Deng F, Xu X, Lv M, et al. Age is associated with prognosis in serous ovarian carcinoma. J Ovarian Res 2017;10:36. [Crossref] [PubMed]
- Murata M, Thanan R, Ma N, et al. Role of nitrative and oxidative DNA damage in inflammation-related carcinogenesis. J Biomed Biotechnol 2012;2012:623019 [Crossref] [PubMed]
- Turnu L, Di Minno A, Porro B, et al. Assessing Free-Radical-Mediated DNA Damage during Cardiac Surgery: 8-Oxo-7,8-dihydro-2'-deoxyguanosine as a Putative Biomarker. Oxid Med Cell Longev 2017;2017:9715898 [Crossref] [PubMed]
- Di Minno A, Turnu L, Porro B, et al. 8-Hydroxy-2-deoxyguanosine levels and heart failure: A systematic review and meta-analysis of the literature. Nutr Metab Cardiovasc Dis 2017;27:201-8. [Crossref] [PubMed]
- Yasuda N, Bolin C, Cardozo-Pelaez F, et al. Effects of repeated bouts of long-duration endurance exercise on muscle and urinary levels of 8-hydroxy-2'-deoxyguanosine in moderately trained cyclists. J Sports Sci 2015;33:1692-701. [Crossref] [PubMed]
- Fogarty MC, Devito G, Hughes CM, et al. Effects of alpha-lipoic acid on mtDNA damage after isolated muscle contractions. Med Sci Sports Exerc 2013;45:1469-77. [Crossref] [PubMed]
- Soares JP, Silva AM, Oliveira MM, et al. Effects of combined physical exercise training on DNA damage and repair capacity: role of oxidative stress changes. Age (Dordr) 2015;37:9799. [Crossref] [PubMed]
- Villaño D, Vilaplana C, Medina S, et al. Effect of elite physical exercise by triathletes on seven catabolites of DNA oxidation. Free Radic Res 2015;49:973-83. [Crossref] [PubMed]
- Sasaki S, Matsuura T, Takahashi R, et al. Effects of regular exercise on neutrophil functions, oxidative stress parameters and antibody responses against 4-hydroxy-2-nonenal adducts in middle aged humans. Exerc Immunol Rev 2013;19:60-71. [PubMed]
- Debelec-Butuner B, Ertunc N, Korkmaz KS. Inflammation contributes to NKX3.1 loss and augments DNA damage but does not alter the DNA damage response via increased SIRT1 expression. J Inflamm (Lond) 2015;12:12. [Crossref] [PubMed]
- Mrakic-Sposta S, Gussoni M, Moretti S, et al. Effects of Mountain Ultra-Marathon Running on ROS Production and Oxidative Damage by Micro-Invasive Analytic Techniques. PLoS One 2015;10:e0141780 [Crossref] [PubMed]
- Poulsen HE, Loft S, Vistisen K. Extreme exercise and oxidative DNA modification. J Sports Sci 1996;14:343-6. [Crossref] [PubMed]
- Pilger A, Germadnik D, Formanek D, et al. Habitual long-distance running does not enhance urinary excretion of 8-hydroxydeoxyguanosine. Eur J Appl Physiol Occup Physiol 1997;75:467-9. [Crossref] [PubMed]
- Kabasakalis A, Tsalis G, Zafrana E, et al. Effects of endurance and high-intensity swimming exercise on the redox status of adolescent male and female swimmers. J Sports Sci 2014;32:747-56. [Crossref] [PubMed]
- Bloomer RJ, Fisher-Wellman KH. Blood oxidative stress biomarkers: influence of sex, exercise training status, and dietary intake. Gend Med 2008;5:218-28. [Crossref] [PubMed]
- Okamura K, Doi T, Hamada K, et al. Effect of repeated exercise on urinary 8-hydroxy-deoxyguanosine excretion in humans. Free Radic Res 1997;26:507-14. [Crossref] [PubMed]
- Radák Z, Pucsuk J, Boros S, et al. Changes in urine 8-hydroxydeoxyguanosine levels of super-marathon runners during a four-day race period. Life Sci 2000;66:1763-7. [Crossref] [PubMed]
- Miyata M, Kasai H, Kawai K, et al. Changes of urinary 8-hydroxydeoxyguanosine levels during a two-day ultramarathon race period in Japanese non-professional runners. Int J Sports Med 2008;29:27-33. [Crossref] [PubMed]
- Almar M, Villa JG, Cuevas MJ, et al. Urinary levels of 8-hydroxydeoxyguanosine as a marker of oxidative damage in road cycling. Free Radic Res 2002;36:247-53. [Crossref] [PubMed]
- Dizdaroglu M, Jaruga P, Rodriguez H. Measurement of 8-hydroxy-2'-deoxyguanosine in DNA by high-performance liquid chromatography-mass spectrometry: comparison with measurement by gas chromatography-mass spectrometry. Nucleic Acids Res 2001;29:E12 [Crossref] [PubMed]
- Kurgan Ş, Onder C, Altingoz SM, et al. High sensitivity detection of salivary 8-hydroxy deoxyguanosine levels in patients with chronic periodontitis. J Periodontal Res 2015;50:766-74. [Crossref] [PubMed]
- Kondo S, Toyokuni S, Tanaka T, et al. Overexpression of the hOGG1 gene and high 8-hydroxy-2'-deoxyguanosine (8-OHdG) lyase activity in human colorectal carcinoma: regulation mechanism of the 8-OHdG level in DNA. Clin Cancer Res 2000;6:1394-400. [PubMed]
- Karpouzi C, Nikolaidis S, Kabasakalis A, et al. Exercise-induced oxidatively damaged DNA in humans: evaluation in plasma or urine? Biomarkers 2016;21:204-7. [Crossref] [PubMed]
- Thompson CM, Fedorov Y, Brown DD, et al. Assessment of Cr(VI)-induced cytotoxicity and genotoxicity using high content analysis. PLoS One 2012;7:e42720 [Crossref] [PubMed]
- Kim J, Kim NH, Sohn E, et al. Methylglyoxal induces cellular damage by increasing argpyrimidine accumulation and oxidative DNA damage in human lens epithelial cells. Biochem Biophys Res Commun 2010;391:346-51. [Crossref] [PubMed]
- Cafueri G, Parodi F, Pistorio A, et al. Endothelial and smooth muscle cells from abdominal aortic aneurysm have increased oxidative stress and telomere attrition. PLoS One 2012;7:e35312 [Crossref] [PubMed]
- Puente BN, Kimura W, Muralidhar SA, et al. The oxygen-rich postnatal environment induces cardiomyocyte cell-cycle arrest through DNA damage response. Cell 2014;157:565-79. [Crossref] [PubMed]
- Tsai-Turton M, Luong BT, Tan Y, et al. Cyclophosphamide-induced apoptosis in COV434 human granulosa cells involves oxidative stress and glutathione depletion. Toxicol Sci 2007;98:216-30. [Crossref] [PubMed]
- Debelec-Butuner B, Bostanci A, Heiserich L, et al. Automated Cell-Based Quantitation of 8-OHdG Damage. Methods Mol Biol 2016;1516:299-308. [Crossref] [PubMed]
- Runge R, Hiemann R, Wendisch M, et al. Fully automated interpretation of ionizing radiation-induced gammaH2AX foci by the novel pattern recognition system AKLIDES(R). Int J Radiat Biol 2012;88:439-47. [Crossref] [PubMed]
- Willitzki A, Lorenz S, Hiemann R, et al. Fully automated analysis of chemically induced gammaH2AX foci in human peripheral blood mononuclear cells by indirect immunofluorescence. Cytometry A 2013;83:1017-26. [Crossref] [PubMed]
- Reddig A, Fatahi M, Roggenbuck D, et al. Impact of in Vivo High-Field-Strength and Ultra-High-Field-Strength MR Imaging on DNA Double-Strand-Break Formation in Human Lymphocytes. Radiology 2017;282:782-9. [Crossref] [PubMed]
- Fatahi M, Reddig A. DNA double-strand breaks and micronuclei in human blood lymphocytes after repeated whole body exposures to 7T Magnetic Resonance Imaging. Neuroimage 2016;133:288-93. [Crossref] [PubMed]
- Reddig A, Fatahi M, Friebe B, et al. Analysis of DNA Double-Strand Breaks and Cytotoxicity after 7 Tesla Magnetic Resonance Imaging of Isolated Human Lymphocytes. PLoS One 2015;10:e0132702 [Crossref] [PubMed]
- Rasche L, Heiserich L, Behrens JR, et al. Analysis of Lymphocytic DNA Damage in Early Multiple Sclerosis by Automated Gamma-H2AX and 53BP1 Foci Detection: A Case Control Study. PLoS One 2016;11:e0147968 [Crossref] [PubMed]
- Leifert WR, Siddiqui SM. gammaH2AX is a biomarker of modulated cytostatic drug resistance. Cytometry A 2015;87:692-5. [Crossref] [PubMed]
- Menegakis A, von Neubeck C, Yaromina A, et al. gammaH2AX assay in ex vivo irradiated tumour specimens: A novel method to determine tumour radiation sensitivity in patient-derived material. Radiother Oncol 2015;116:473-9. [Crossref] [PubMed]
- Reddig A, Lorenz S, Hiemann R, et al. Assessment of modulated cytostatic drug resistance by automated gammaH2AX analysis. Cytometry A 2015;87:724-32. [Crossref] [PubMed]
- European Standards Committee on Urinary (DNA) Lesion Analysis. Toward consensus in the analysis of urinary 8-oxo-7,8-dihydro-2'-deoxyguanosine as a noninvasive biomarker of oxidative stress. FASEB J 2010;24:1249-60. [Crossref] [PubMed]
Cite this article as: Korkmaz KS, Debelec Butuner B, Roggenbuck D. Detection of 8-OHdG as a diagnostic biomarker. J Lab Precis Med 2018;3:95.


